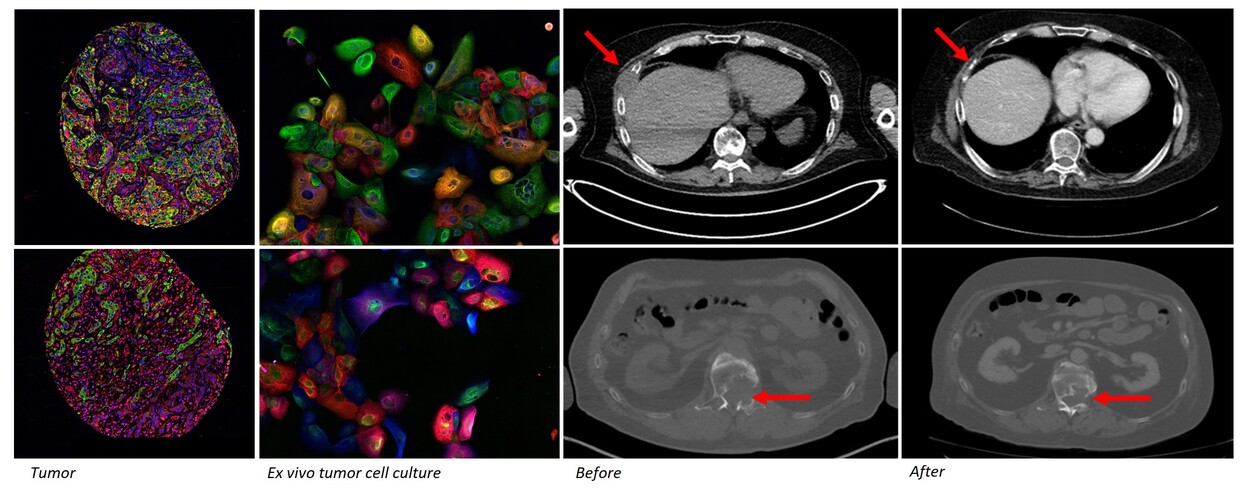
Currently, all stages of drug discovery and basic molecular cancer research rely on established cancer cell lines. The most comprehensively utilized collection of established cancer cell lines, composing the CCLE collection and underlying e.g., the DepMap data, consist of around 2000 cancer cell lines. The median year of establishment of these cell lines is 1991, pre-dating development of targeted cancer therapies and spanning all the way back to the 1950s. Many key cellular properties such lineage heterogeneity, genomic stability and mutational burden change during prolonged culture of cancer cell lines and consequently, the discovery and research of novel therapeutic targets and analysis of e.g., drug resistance mechanism can be significantly biased. Novel cancer cell lines and tumor models representing patients treated with the current cancer therapies are therefore urgently needed to allow more clinically relevant research of the efficacy and resistance mechanism to the latest novel cancer therapies.
21st century cancer research requires 21st century model cell lines
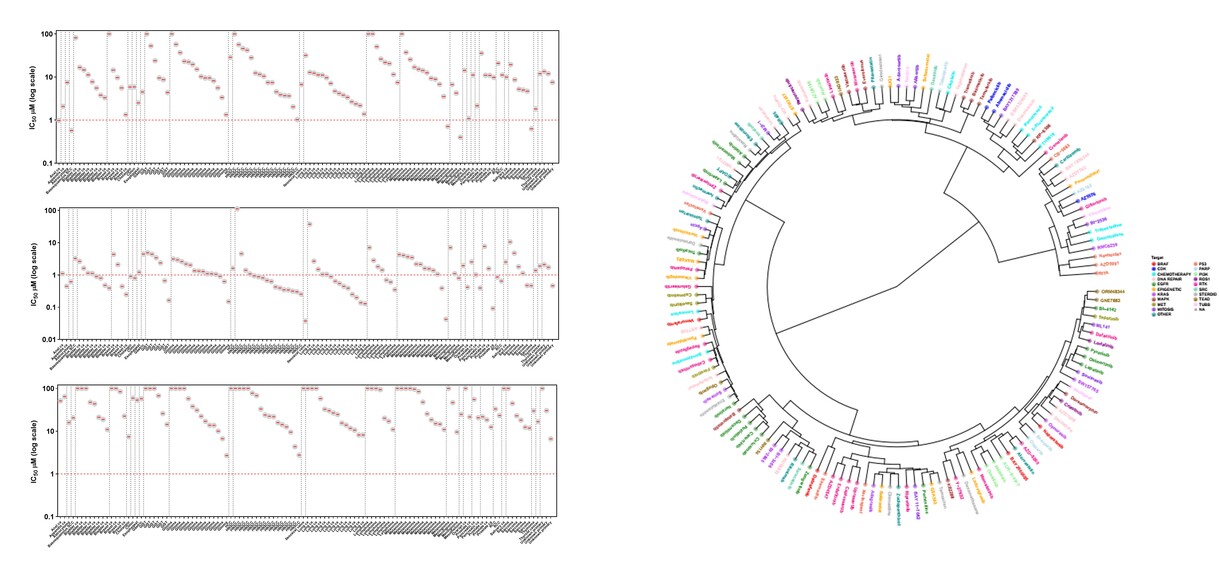
In context of our ongoing clinical research activities, we are building a comprehensive collection of new cancer model cell lines to serve the needs of cancer research and drug development. In parallel to collection of tumor samples for ex vivo drug efficacy testing, we´ve established a proprietary cell line repository consisting of 150+ cell lines representing 50+ different solid cancer typess (@01/2026); the MISB Cancer Cell Line Panel. The continuously expanding MISB cell line panel has been characterized extensively in context of Misvik’s Precision Oncology initiative including using DNA and RNA sequencing and extensive drug screens of hundreds of cancer therapies, including the very latest novel cancer drugs. The MISB cell line panel offers a collection of low-passage clinically relevant tumor cell models that retain the original patient-specific tumor biology including lineage heterogeneity, genetic background and treatment history (62 different therapies covered as previous line of treatment before relapse and establishment of cell line). The MISB cell line panel is offered for rapid large scale high-throughput cell panel screening, in-depth studies of drug efficacy and combinatorial synergy as well as for genome-wide loss of function genomic screening using siRNA and CRISPR.
Common and rare cancers
The common cancer types included in the current MISB cell line panel (@01/2026) include: Bladder, Breast, Cholangiocarcinoma, Colon, Esophageal, Endometrial, Glioma, Head and Neck, Lung, Melanoma, Ovarian, Pancreatic, Prostate and Renal. Rare cancer types include: Conjunctival melanoma, Choroidal melanoma, DSRCT, GIST, Ileocecal valve, Large Cell Neuroendocrine PCa, Salivary gland, Sarcomas, Urachal and cancers of unknown origin.
Pan-cancer including first from indication
Diverse set of common and rare solid cancers covering 50+ different cancer types
Extensive characterization
Detailed molecular and clinical profiling including DNA & RNAseq, proteomics and large-scale drug sensitivity profile
Targeted therapy relevance
Established from tumors of heavily pretreated patients covering 64 targeted therapies
Proprietary
No license fees = competitive service pricing and freedom to operate
Patient derived functional tumor models retain the cellular composition and differentiation state heterogeneity of the parental tumors. Primary ex vivo models also reflect the pathology and clinical behavior of the cancer. Misvik´s ex vivo tumor models are a continuously expanding collection of early passage patient derived functional tumor models cryopreserved following initial tissue dissociation and establishment of organoid culture. The ex vivo model collection currently contains 700+ tumor samples representing 90+ different solid cancer types including rare to ultra rare cancers. The sample series also contains matched longitudinal samples from the same patient prior and after treatment allowing study of therapy efficacy in treatment naive and relapsed tumor models. All samples are incorporated on Misvik´s master RPPA (reverse-phase-protein array) sample repository for rapid target evaluation and general protein expression analysis across different cancers types.
Building insights on latest novel cancer drugs using pan-cancer ex vivo tumor models.
Biologically relevant ex vivo tumor models are primary cultures isolated directly from dissociated tumor samples